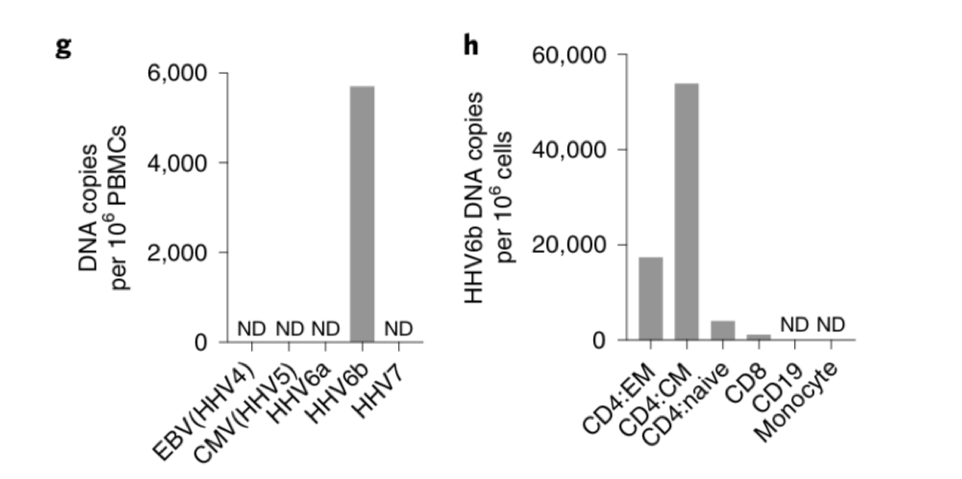
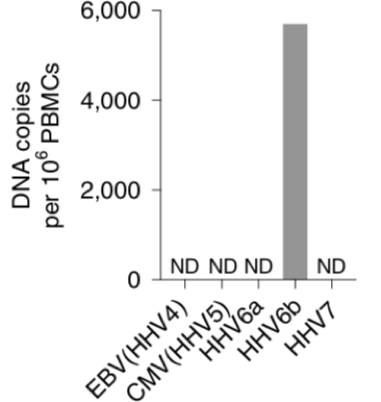
Researchers at the NIH used RNA-Seq cells from skin and blood to study the underlying mechanisms in DIHS/DRESS and identified both HHV-6 and JAK inhibitors as potential targets. They also found central memory CD4+T cells enriched with HHV-6B.
 The group from the Microbiome and Inflammation Section, Dermatology Branch, of the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS), NIH, used single-cell RNA-Seq to study a case of refractory DIHS/DRESS in a 44 year old patient. The group hypothesized that single-cell RNA sequencing (scRNA-seq) could be used to identify overexpressed genes or altered pathways that could be targeted by novel therapies. After finding high levels of HHV-6B in central memory CD4 T-cells, and identifying the JAK-STAT signaling pathway as a potential target, they treated the patient successfully with a combination of a JAK inhibitor and the antiviral drug ganciclovir.
The group from the Microbiome and Inflammation Section, Dermatology Branch, of the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS), NIH, used single-cell RNA-Seq to study a case of refractory DIHS/DRESS in a 44 year old patient. The group hypothesized that single-cell RNA sequencing (scRNA-seq) could be used to identify overexpressed genes or altered pathways that could be targeted by novel therapies. After finding high levels of HHV-6B in central memory CD4 T-cells, and identifying the JAK-STAT signaling pathway as a potential target, they treated the patient successfully with a combination of a JAK inhibitor and the antiviral drug ganciclovir.
The group determined that the patient's T cells in both skin and blood exhibited increased proliferation, distinct chemokine receptor expression, and upregulated genes involved in the JAK–STAT signaling pathway. HHV6-B was primarily enriched in circulating CD4+ T cells with a memory cell phenotype.
While many publications have shown that viral reactivation occurs in most DIHS/DRESS cases, the contribution of HHV-6 to its pathogenesis remains controversial.
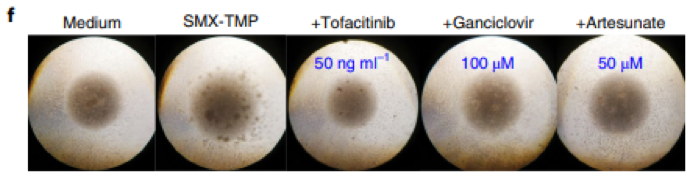
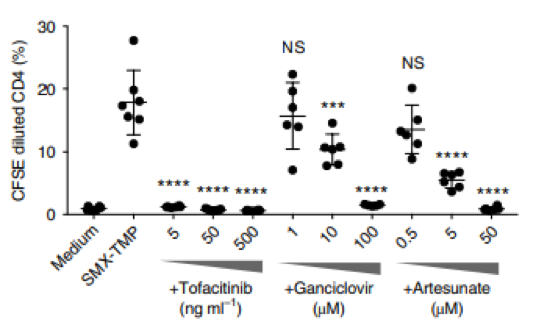
 The authors used the lymphocyte transformation test (LTT) to further model the possible capabilities of their newly identified therapeutic targets. When they tested ganciclovir and artesunate (both known to have anti-HHV-6 activity) in their LTT model, they discovered that both significantly suppressed CD4+ T cell proliferation. The authors conclude that while early events in the disease and the contribution of viruses remain to be elucidated, LTT studies suggested that both JAK inhibition and antiviral agents are promising targets in drug-induced phases of DiHS/DRESS.
The authors used the lymphocyte transformation test (LTT) to further model the possible capabilities of their newly identified therapeutic targets. When they tested ganciclovir and artesunate (both known to have anti-HHV-6 activity) in their LTT model, they discovered that both significantly suppressed CD4+ T cell proliferation. The authors conclude that while early events in the disease and the contribution of viruses remain to be elucidated, LTT studies suggested that both JAK inhibition and antiviral agents are promising targets in drug-induced phases of DiHS/DRESS.
Read the full paper: Kim 2020